
informatik



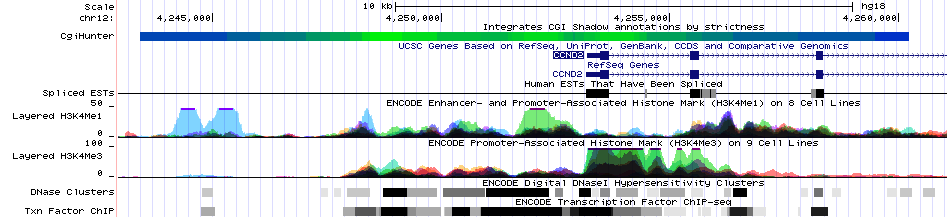 |
Visualisation of the CpG Mountain annotation in the UCSC Genome Browser Green indicates areas with strong CpG island characteristics (CGI cores) Blue indicates areas with weaker characteristics (CGI shores) |
|
CpG islands are genome regions with an elevated Guanine + Cytosine content (GC content) and a higher CpG dinucleotide frequency than observed in the bulk genome. More than half of the human gene promoters colocalize with CpG islands and their methylation status has been shown to correlate with the expression level of the associated genes. As a consequence, CpG islands are assumed to be hotspots of epigenetic regulation.
However, an exact definition of CpG islands is still not agreed upon.
The CgiHunter software tool follows the original sequence-based CpG island definition proposed by Gardiner-Garden and Frommer.
They defined three requirements a genome region has to fulfill to qualify as a CpG island: a minimal length, a minimal GC content and a minimal ratio of its observed over its expected CpG dinucleotide frequency.
Unlike many other heuristic-based approaches, the CgiHunter algorithm has been proven to identify all genome regions that meet a specified criteria and results in robust and consistent CpG island annotations.
CgiHunter allows a free choice the three parameters to allow customizibility of annotations.
We applied CgiHunter to generate 100 annotations using a grid of different parameters. Subsequently, we applied publical available data on epigenetic marks, such as absence of DNA methylathin, promoter marking histone modifications and overlap with RNA polymerase II occupancy sites, to integrate these 100 CpG island annotations into a single heatmap - the CpG Mountain annotation. Currently, the paper that describes the method in detail, is under revision. If you have questions regarding the underlying algorithm, please contact us. Annotations for the human genome builds hg18 and hg19 are available on this website. The latest version of software is also available on this website. If you have further questions, do not hesitate to contact us (CgiHunter@mpi-inf.mpg.de) or visit our FAQ page. |